Congratulations to Clay Klein for Receiving the 2025 Nick Cobb Memorial Scholarship from SPIE
STROBE and JILA graduate student Clay Klein has been awarded the prestigious 2025 Nick Cobb Memorial Scholarship, presented by SPIE, the International Society for Optics and Photonics, and Siemens EDA. The scholarship, valued at $10,000, recognizes Klein’s outstanding contributions to the field of optics and photonics.
“I am honored to be awarded the Nick Cobb Memorial Scholarship,” Klein stated. “This scholarship provides me with the exciting opportunity to share my research in this field and connect with others in the industry at the SPIE conference in February.”
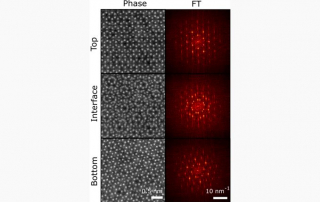
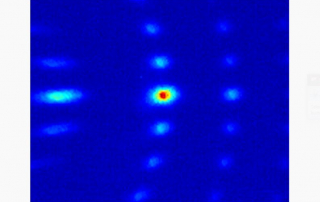
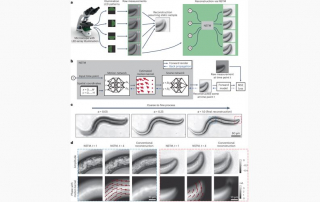
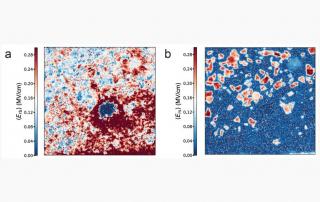
Klein conducts research in the laboratories of JILA Fellows and University of Colorado Boulder Physics professors Margaret Murnane and Henry Kapteyn. His work focuses on cutting-edge advancements in nanoscale extreme ultraviolet imaging science.
The award will be formally presented during the Welcome and Plenary Presentation at the SPIE Advanced Lithography + Patterning Conference in San Jose, California, on February 24, 2025. Congratulations Clay!